In recent years, several NGS programs have sequenced large number of patients with different types of neuromuscular diseases including limb girdle muscular dystrophies, congenital myasthenic syndromes and inherited neuropathies, among others. Most of these studies, such as MYO-SEQ have been performed in Caucasic populations of European origin or other groups descendent of European ancestors. These studies have informed us about the epidemiology of these diseases in Europe, have provided diagnosis to a large number of patients and have also allowed the identification of novel genes causing neuromuscular diseases. However, little is known regarding the prevalence of common neuromuscular diseases in other ethnicities, such as the Amerindians which are predominant in most of the Andean countries of South and Central America.
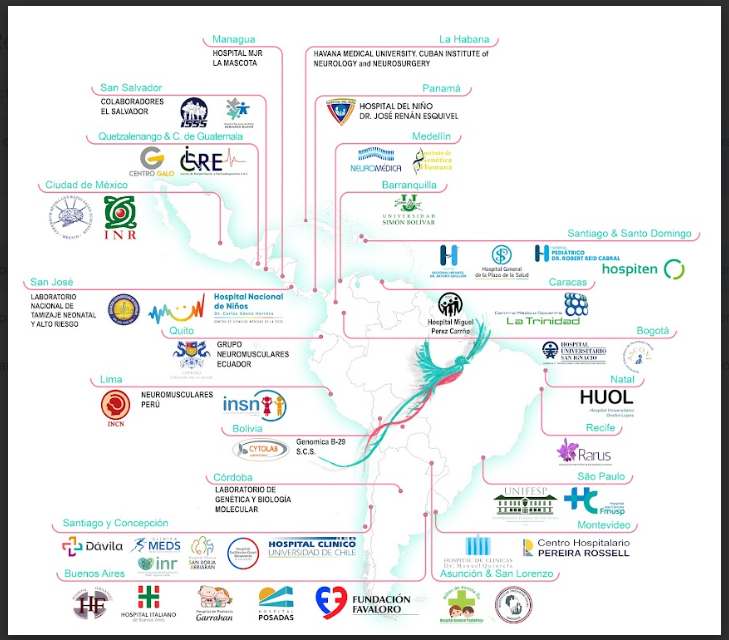
We aim to develop a large multinational multicentric diagnostic study using next generation sequencing in patients with neuromuscular diseases who are followed up in highly specialised units in Latin America. The implementation of a research projects using NGS technologies applied to DNA samples of NMD patients from these naïve areas can provide valuable epidemiological information, but will also contribute to increase the number of patients genetically diagnosed and in turn to identify patients to be included in the new upcoming clinical trials or those who could benefit of the new treatments that are being approved and required a final undoubtful diagnosis. Moreover, these studies could identify new candidate genes and diseases; this will also result in additional diagnoses for patients in those countries but also around the world.

Ana Topf
Senior Research Associate

Alejandro Gonzalez Chamorro
Project Manager

Jordi Díaz Manera
Professor of Neuromuscular Disease